Introduction to Data Analysis with R
Hello and welcome!
… to the latest iteration of my introductory R course, where we will learn to analyse data in style.
Current course dates
SS25
2025-04-14 – 2025-05-19
See structure of the course
Lecture online in your own time
Seminar twice weekly on Monday and Thursday at 13:00
In the Mathematikon (INF 205),
Seminar room 11, 5.OG (Mondays)
and PC-Pool 2 Raum 3/104 (Thursdays)
Sign-up (Heidelberg University students): see discord
Language: Lectures are in English but the seminar can be in German if you choose so (in any case, you can always ask questions in German as well).
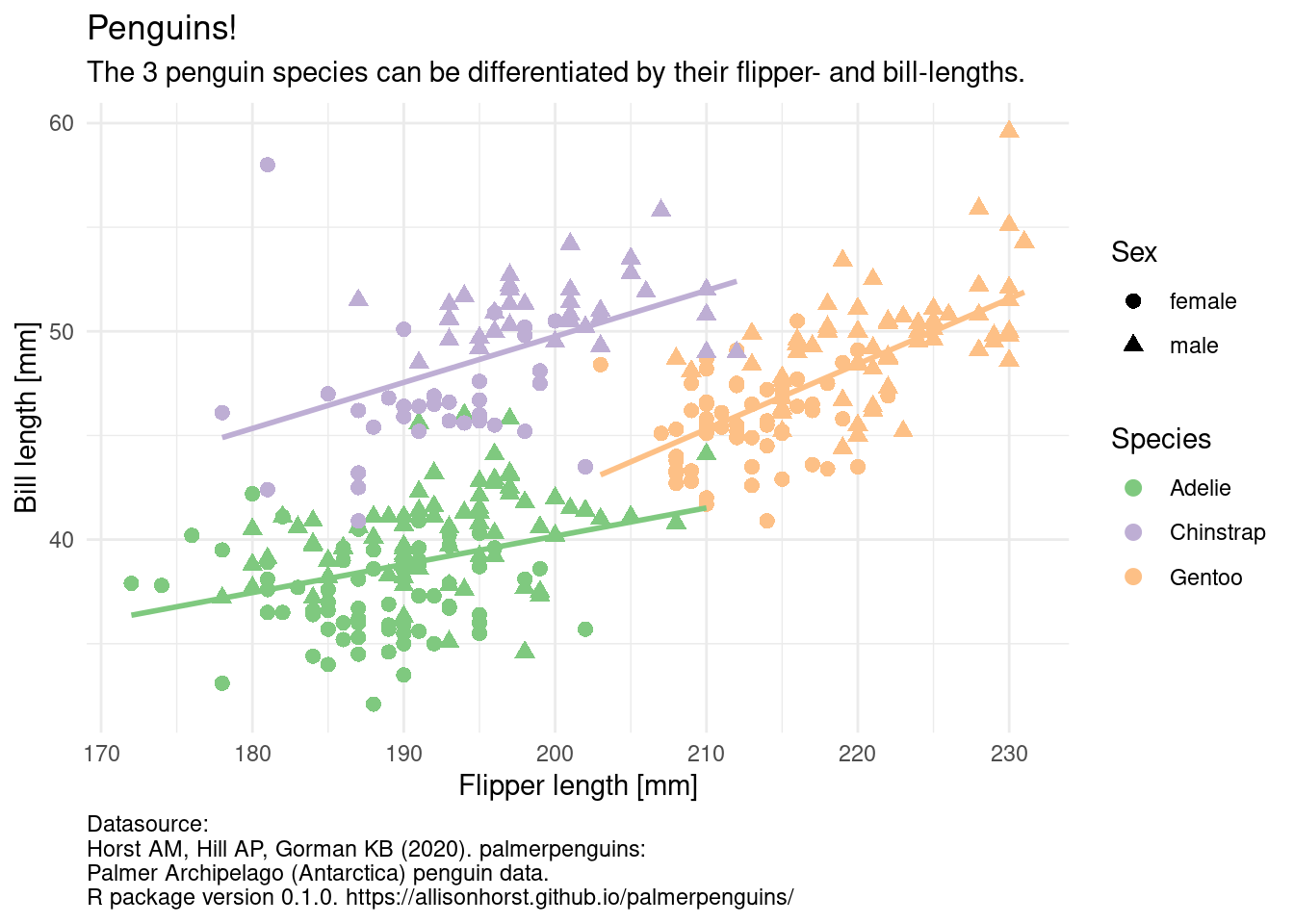
In this course, we will handle different kinds of data, create pretty and insightful visualizations, compute different statistics on our data and also explore what these statistical concepts mean. From penguins to p-values, I got you covered.
Prerequisits
No prior knowledge necessary.
Software to install:
Structure of the course
Most participants will be biochemistry bachelor (and master) students, but the material is open to anyone!
- There are 8 lectures in total, each accompanied by:
- A video of the lecture at the top of each page
- The lecture script, which consists of the code written during the lecture (plus some more code to generate illustrative graphics) and explanations
- Exercises to complete and hand in
- A seminar to discuss the exercises
- A discord server to ask questions and share solutions
In addition to the 8 regular lectures + follow up seminars, there is an introductory seminar in the first week, before watching the first lecture.
Key dates:
- April 14 (Monday, 13:00): First seminar
- April 17 (Thursday, 13:00): Seminar for the first lecture
- May 19: Seminar for the last lecture (Note, that this is 1 week later than would be expected according to a perfect fit of 2 lectures per week, because there are two public holidays (April 21 & May 1) in the timespan that are excluded)
The first seminar is dedicated to helping you get your laptops set up for the course, show the course structure and explain core concepts.
I do recommend to watch the lecture in your own time, and then use the lecture script afterwards to look up concepts and code you want to revisit. Code chunks also have a copy-button, which is helpful for quickly playing around with it, but make sure you actually walk through the lecture and do the typing first, because muscle memory will server you well in the future.
Exercises
To complete the course, hand in at least 7 out of 8 exercises. The important part here is not that each exercise is a perfect solution, but if you encounter questions and struggles during your attempt of the exercise make sure to include those pain points as well, so that we can cover those in the Seminar. Please hand in your solutions before the seminar via a direct message on discord. The earlier you submit your solutions, the more time I have to prepare specific answers to your questions for the seminar.
Seminar
Each week, we will meet to discuss the exercises and answer any questions that might have have popped up. We will meet in:
Mathematikon (INF 205)
Seminar room 11, 5.OG (Mondays)
PC-Pool 2 Raum 3/104 (Thursdays)
Please bring your own laptop, so that you can code along and know that you will be able to apply what you learned after the course as well.
Discord and signup
If you are a biochemistry student at Heidelberg University, click on this link: https://discord.gg/TYQhFAAfxu to join our discord server to sign up for the course. On the server, you will be able to ask questions that can be answered by me and your fellow learners, hand in the exercises and receive feedback.
I will also need your full name and matriculation number, so that we can put the course onto your official transcript of records after completion.